|
|
Nucleic acid structure and function
- Description
- DNA = deoxy-ribo-nucleic acid
- RNA = ribo-nucleic acid
- deoxy = 2'-H (vs 2'-OH in RNA)
- ribo = ribose sugar = pentose sugar
- acid = phosphate group gives it acidity
- Nucleotides and nucleosides
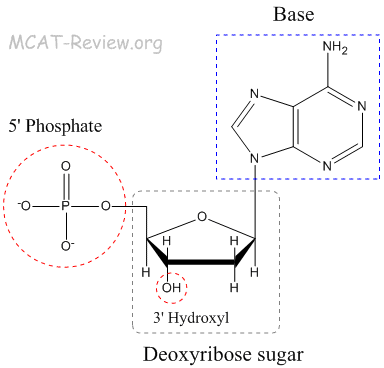
- Nucleotide = base (Adenine, Guanine, Thymine, Cytosine) + sugar + phosphate.
- Nucleoside = base + sugar = Adenosine, Guanosine, Thymidine, Cytidine.
- Sugar phosphate backbone = phosphodiester bonds linking the 5' phosphate to the 3' hydroxyl group of the pentose sugar
- Base can either be purines A and G (the big ones with 2 rings) or pyrimidines T and C (the small ones with 1 ring).
- Thymine is replaced with Uracil in RNA
- The phosphate group gives DNA its acidity.
- Deoxyribonucleic acid (DNA): double helix, Watson-Crick model of DNA structure
- The "double" in the double helix means that DNA is found in a double-stranded form - 2 single-stranded chains of DNA stuck to each other via hydrogen bonding of the base pairs.
- The 2 single-strands are anti-parallel to each other. Going from 5' to 3' of one strand means going from 3' to 5' of the other strand.
- The "helix" in the double helix means that the entire thing is wound up in a spiral.
- Base pairing specificity: A with T, G with C
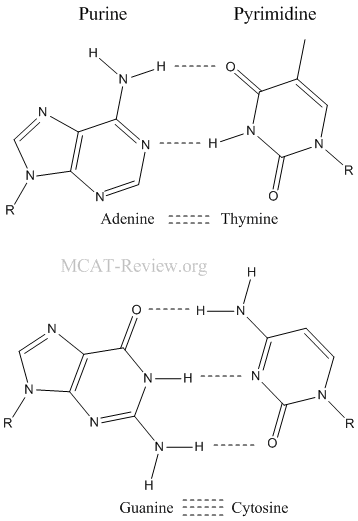
- A forms 2 hydrogen bonds with T.
- G forms 3 hydrogen bonds with C.
- GC bonds are stronger. DNA with high GC content will be harder to break apart.
- Complementary strands of DNA hydrogen bond with each other.
- 5'-ATGC-3' will be complementary to 5'-GCAT-3' or 3'-TACG-5', but NOT 5'-TACG-3'. make sure you get the 5's and 3's right.
- Function in transmission of genetic information
- Because of the complementary nature of base pairing, DNA can transmit genetic information through replication.
- DNA denaturation, reannealing, hybridization
- denaturation = separation of 2 complementary strands = caused by high temperature
- reannealing = coming together of the 2 complementary strands after they've been separated
- hybridization = annealing = coming together of complementary strands
- annealing is generally used in PCR reactions, where primer anneals to the template strand
- hybridization is generally used in Southern blotting (probe anneals to target) and plasmid ligation (sticky ends anneal)
DNA replication
- Mechanism of replication: separation of strands, specific coupling of free nucleic acids
- First, the double stranded DNA must separate, or unwind. To do this:
- DNA gyrase (class II topoisomerase) is responsible for uncoiling the DNA ahead of the replication fork.
- Helicase is responsible for unwinding the DNA at the replication fork.
- Single-strand binding protein (SSB) is responsible for keeping the DNA unwound after the helicase. SSBs stabilize single-stranded DNA by binding to it.
- Next, you start making DNA that is complementary to the newly unwound/separated DNA. Note, all biological DNA synthesis occurs from the 5' to the 3' end.
- Primase gets this started by laying down a short RNA primer on the unwound DNA. The primer is made of RNA, but is complementary to the DNA sequence. Later, this RNA is replaced with DNA.
- DNA polymerase then takes over and makes DNA that is complementary to the unwound DNA.
- DNA synthesis occurs on both strands of the unwound DNA. The synthesis that proceeds in the direction of the replication fork is the leading strand. The synthesis that proceeds in the opposite direction to the replication fork is the lagging strand. The lagging strand contains Okazaki fragments.
- Finally, RNA primers are replaced with DNA by a special DNA polymerase. The Okazaki fragments in the lagging strands are then stitched together by DNA ligase.
- DNA synthesis is bidirectional: 2 replication forks form and proceeds in opposite directions (like an expanding bubble).
- Biological DNA synthesis always proceeds from the 5' end to the 3' end.
- DNA polymerase has proof-reading activity, which means it corrects any mistakes (mutations) it makes.
- Replication occurs once every cell generation, during the S phase. (Cell division may occur twice in meiosis, but replication still occurs once only)
- Semi-conservative nature of replication
- Newly synthesized DNA contains one old strand and one new strand.
- Meselson and Stahl proved this by experiment: Basically, they used heavy (15N) DNA as the old (pre-replication) DNA, and used light (14N) nucleotides for the synthesis of new DNA. They can tell the difference between heavy and light DNA by centrifugation. What they found was that when heavy DNA undergoes one round of replication in light nucleotides, the DNA made is of intermediate weight. After the second round of replication, the DNA is split between intermediate and light weight.
- If DNA replication were completely conservative, only heavy and light DNA would be seen, and nothing in between. This was not the case.
- If DNA replication were dispersive, everything would be of intermediate weight. Again, this was not the case because after the second round of replication, light DNA was seen.
Repair of DNA
- Repair during replication
- DNA polymerase has proof-reading activity (also called 3' → 5' exonuclease activity). If a wrong nucleotide gets incorporated, the polymerase will "back-up" and replace it with the correct one.
- The special polymerase that replaces the RNA primers with DNA also have 5' → 3' activity. This allows the polymerase to clear away short stretches of incorrect nucleotides (RNA or incorrect DNA) and replace it with the right ones (DNA). This process is also called repair.
- Repair of mutations
- Mismatch repair: enzymes recognize incorrectly paired base-pairs and cuts out the stretch of DNA containing the mismatch. Then polymerase re-adds the correct nucleotides in.
- During mismatch repair, the repair enzyme must decide what strand of DNA to cut since DNA contains 2 strands. To do this, the enzyme cuts the DNA strand that do not have methylations. The original (old) DNA has methylations, but the newly synthesized DNA do not have them until shortly after replication. Thus, there is a window of time when mismatch repair enzymes can know what strand to cut if mismatch is encountered.
- Base-excision repair: a damaged base gets cut out. Then the base's sugar phosphate backbone gets cut out. And then, several more nucleotides next to the base get cut out. Finally, polymerase remakes the cut out nucleotides.
- Nucleotide-excision repair: damaged nucleotide(s) gets cut out and then polymerase replaces it. This is like mismatch-repair, but it's not for mismatch. It's for damages like thymine dimers, and other damages that changes normal nucleotides into abnormal nucleotides.
- Nick translation: this is basically 5' → 3' exonuclease activity coupled to polymerase activity. The polymerase here chugs along, chews off the bad nucleotides and then replaces them with new nucleotides. This is what happens when RNA primers are replaced with DNA.
- SOS response in E. Coli: during replication, when there's just too much DNA damage for normal repair to handle, the SOS repair system comes along. Instead of correcting any DNA damages during replication, the polymerase replicates over the damaged DNA as if it were normal. By using the damaged DNA as a template error rates are high, but it's still better than not replicating at all.
Recombinant DNA
- Gene cloning
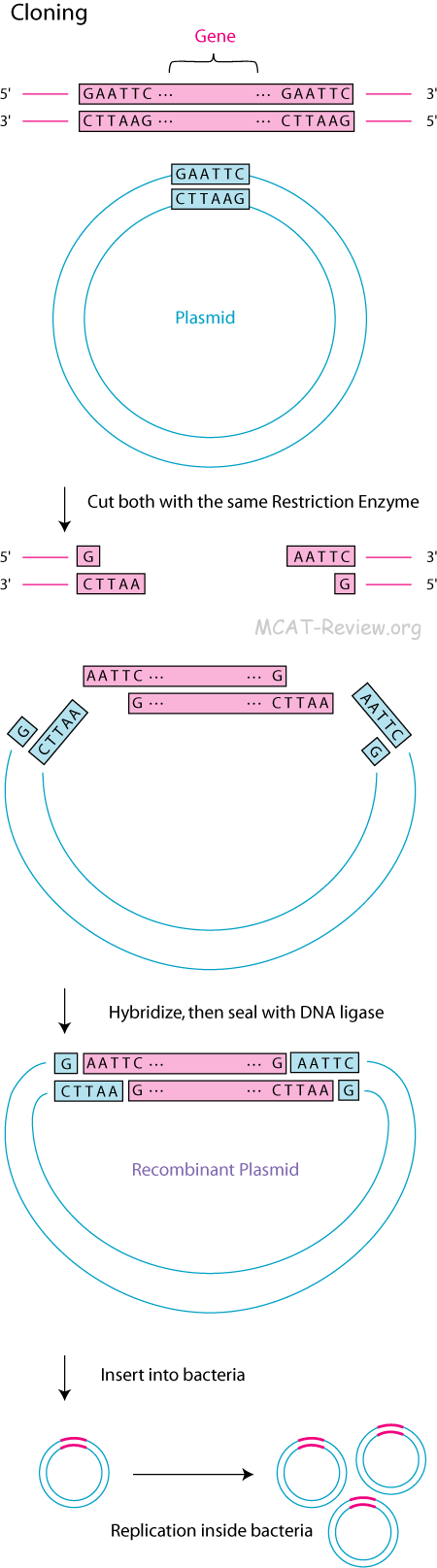
- The plasmid must have a restriction site because you need to open it up for the insertion of your gene.
- The plasmid must have an origin of replication because you want to clone your gene, which is inside your plasmid.
- The plasmid must have an antibiotic resistant gene because this lets you kill competing, useless bacteria that doesn't have your plasmid. When you add an antibiotic, only the bacteria with the antibiotic resistant plasmid will live.
- Plasmids replicate independently of the genomic DNA of the bacteria.
- Restriction enzymes
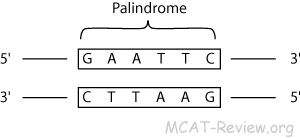
- Restriction enzymes (also called restriction endonucleases) cut double stranded DNA at palindrome sequences. The resulting fragments are called restriction fragments.
- If you read from 5' → 3' of one strand, then read from 5' → 3' of the other strand, and they are the same, then the section of the double stranded DNA that you just read is a palindrome sequence.
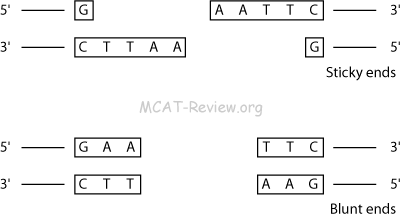
- Some restriction enzymes cut to make sticky ends, which can hybridize.
- Some restriction enzymes cut to make blunt ends, which cannot hybridize.
- DNA libraries = the result of cloning an organism's entire DNA (genomic or cDNA)
- Genomic library = entire genomic DNA -> fragmented by restriction enzyme -> cloned
- cDNA library = entire mRNA population -> reverse transcription into cDNA -> cloned
- Generation of cDNA: mRNA + reverse transcriptase = cDNA
- Hybridization
- Hybridization, also called annealing, is where DNA strands base pair with each other.
- In Southern blotting, DNA probes are used to hybridize onto DNA fragments containing a target sequence.
- In gene cloning, hybridization refers to the process where sticky ends from a restriction fragment of a gene base pairs with the same sticky ends on a plasmid.
- Expressing cloned genes
- Expression = Cloned gene -> mRNA -> protein
- Expression needs a vector with the transcription and translation machinery
- The cloned gene needs to be placed downstream of a strong promoter so that a lot of mRNA will be transcribed
- PCR
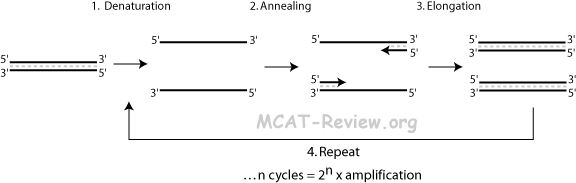
- Denaturation: heat (90 °C) to separate double stranded DNA template.
- Annealing: cool reaction in order for primers to anneal to the now single stranded DNA template.
- Excess amount of primers, so they out complete re-annealing of the template strands.
- Elongation: use heat stable polymerase to extend the primers.
- Repeat steps 1 to 3 for n cycles. The resulting amplification of the original DNA template after n cycles is 2n.
- Gel electrophoresis and Southern blotting
- Gel electrophoresis = separates DNA fragments on a sheet of gel
- Southern blotting = probes make your DNA light up on that sheet of gel
- Probe for southern blotting = DNA that is complementary to the target DNA
- Other blots
- Northern blot: RNA target, DNA/RNA probe
- Western blot: protein target, antibody probe
- DNA sequencing
- Sanger sequencing
- basis: differential chain termination (using dideoxy-NTPs), then detected by capillary electrophoresis
- pros: cheap
- cons: can only sequence a short distance, can only detect mutations if it's at high levels
- Nextgen sequencing = massive parallel sequencing
- basis: addition of different nucleotide causes different color fluoresence, detected on a 2D chip containing many simultaneous reactions
- pros: can sequence an entire genome, can detect mutations at very low levels
- cons: expensive
- Analyzing gene expression
- Gene expression correlates with mRNA levels, so measure RNA
- What genes are expressed and at what levels (global picture)? Use gene expression profiling (microarray). Eg: what genes are overexpressed in certain cancer
- How much is this gene expressed? Use reverse transcription then real time quantitative PCR. Eg: residual BCR-ABL in CML
- Stem cells
- Stem cells = cells that have the potential to differentiate into many different cell types
- Example: hematopoietic stem cells -> neutrophils, lymphocytes, red blood cells, platelets
- Uses: hematopoietic stem cell (bone marrow) transplant as a cure for leukemia
- Practical applications of DNA technology
- Medical applications: PCR coupled with DNA sequencing = detects mutations / variants = used for diagnosis, monitoring disease level, or predict whether a patient will respond to a certain treatment
- Human gene therapy = using viruses to infect and introduce functional genes into human cells
- Pharmaceuticals: uses cloning, expression, and protein purification to isolate proteins/enzymes to treat patients who are deficient in those
- Forensic evidence: DNA fingerprinting = paternity test = identifies a criminal/person
- Environmental cleanup: they are making genetically engineered bacteria that can breakdown oil spills or plastic polymers
- Agriculture: genetically modified foods = introducing genes in crops to make them grow better and be resistant to pests
- Safety and ethics of DNA technology
- Side effect of human gene therapy: the viruses make mistakes when introducing the genes, resulting in mutations that cause cancer
- We are not absolutely certain about the long term health effects of genetically modified foods
- Sequencing someone's entire genome exposes all their underlying genetic diseases - potential basis for discrimination from insurance companies
|
|
